mappa Analysis Pipeline
Version 1.0
Overview
The ICELL8 system allows for a high-throughput, nanowell-based method to isolate nuclei or single cells of any size-and the flexibility to analyze multiple parameters in single experiments. It also uses the power of imaging to distinguish true single cells from multiple cells and live cells from dead cells-allowing you to draw meaningful conclusions from the data.
As automation instruments become more time and cost efficient, it becomes vital to have software that can transform that information into a readily usable format for the scientist. Our easy-to-use solutions are designed for users with all levels of bioinformatics experience-from those analyzing their first sequencing dataset to experts in the field.
mappa™ Analysis Pipeline takes input data from sequencing and the well-list file generated by the CellSelect Software and outputs an HTML report, with results typical to single-cell analysis, and other files, such as a gene matrix, to continue further analysis. R data objects are also output, which are used as input for hanta R kit.
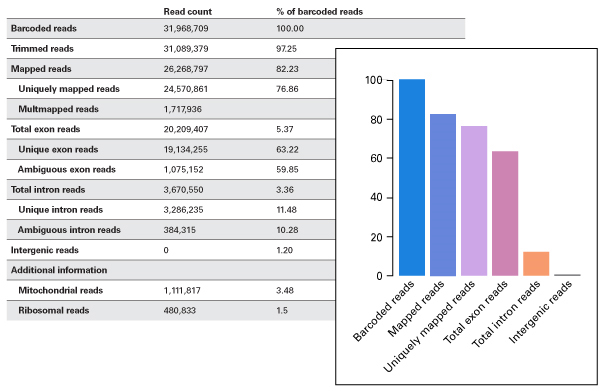 Example output available in the mappa HTML report.
Supported operating systems
Example output available in the mappa HTML report.
Supported operating systems
mappa can be installed on the following versions of Linux:
Hardware requirements
For analyzing Illumina NextSeq
® High-Output data:
- 24-core CPU
- 64 GB+ RAM
- 500 GB+ available hard drive space
Analyzing MiniSeq™ or MiSeq
® data may be possible on servers with less computational power than the specifications above. Analysis of HiSeq
® or NovaSeq™ data would require better hardware than is listed.
Additional third-party software dependencies*
*Not included with the software; must be downloaded and installed separately.
Additional product information
For use in conjunction with workflows on the ICELL8 cx Single-Cell System or ICELL8 Single-Cell System. More information is available in the
mappa Analysis Pipeline User Guide.
mappa software 사용을 원하시면 아래의 링크를 클릭하시어, End User License Agreement를 위해 양식을 작성해 주십시요.
mappa software download




